The honey bee genome has two unusual properties. First – it has the highest recombination rate in animals: Recombination shuffles the genetic deck by mixing up the chromosomes inherited from parents into offspring. Recombination makes the actions of natural selection more efficient because it allows beneficial mutations to spread irregardless of the effects of nearby mutations (see Fig 1).
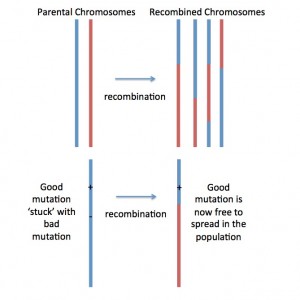
Fig. 1. Recombination shuffles ‘parental chromosomes’ (red and blue), generating ‘mosaic’ or recombined chromosomes. This is how recombination makes natural selection act more effectively: Lets imagine a new beneficial mutation occurring nearby a deleterious (=bad) mutation on the blue chromosome. With no (or low) recombination, the two mutations would be stuck together… forever… and this would prevent natural selection from removing the bad mutation and spreading the good mutation. But if recombination generates a new mosaic chromosome that has the good mutation but not the bad mutation (as pictured), then the good mutation can spread by natural selection!
Second – the bee’s genome has areas that are very rich in the DNA bases G and C, while other areas are very rich in the bases A and T.
Our paper showed that these high GC and low AT genomic regions are maintained by differences in the recombination rate across the bee’s genome: areas with low recombination move towards high AT and areas with high recombination move toward high GC – because of a phenomenon called GC-biased gene conversion. This biased gene conversion is a side effect of high recombination and it acts by increasing the frequency of G or C mutations over A or T mutations.
We also found that regions of the bee genome with high rates of recombination and high GC content have more genetic diversity and evolve a lot faster than genomic regions with low recombination and high AT content.
Now, here comes the fun part… what kind of genes ‘live’ in these high GC genomic areas that have high rates of evolution ? We first looked at high GC genes and examined their biological function in other organisms – turns out many were involved in brain function and things like learning and memory. This is interesting because we know that worker bees (and workers in other social insects) have some of the coolest behaviours out there (remember… they dance!). So we went back to the literature and found several lists of genes that get turned ‘on’ in the brains of workers relative to the brains of queens and drones [
Check out Drones are from Mars, Workers are from Venus!]. It turns out that worker-associated genes are predominantly GC rich, while queen- and drone- associated genes are predominantly AT rich.
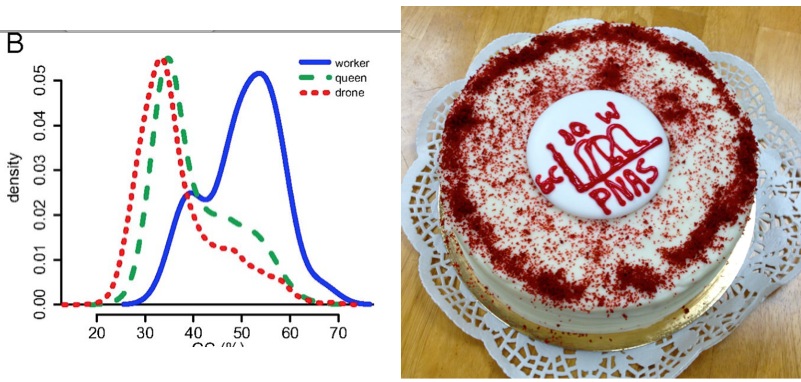
Worker ‘genes’ are GC rich baby! Fig 2 from our PNAS paper, in both tiff and cake formats. The Tiff format is more impactful but the cake format is tastier
So, our results show that the bee’s high recombination rate increases the evolutionary rate of genes associated with worker behaviour, which is an important finding because worker behaviour plays a major role in determining the fitness of insect colonies.
The paper was authored by postdoctoral fellow Dr. Clement Kent, and former Masters students Shermineh Minaei and Brock Harpur.
[edit March 2013]. See Faculty of 1000 recombination, comment by Hunt et al and our reply, and addendum in Communicative and Integrative Biology

